Publications
*co-first authors; #corresponding authors
M. Zhang*, X. Pan*, W. Jung*, A. Halpern, S. W. Eichhorn, Z. Lei, L. Cohen, K. A. Smith, B. Tasic, Z. Yao, H. Zeng, X. Zhuang#. Molecularly defined and spatially resolved cell atlas of the whole mouse brain. Nature 624, 343-354 (2023). [Full text]
Z. Yao#, C. T. J. van Velthoven, M. Kunst, M. Zhang, D. McMillen, … X. Zhuang, H.Zeng#. A high-resolution transcriptomic and spatial atlas of cell types in the whole mouse brain. Nature 624, 317-332 (2023). [Full text]
R. Fang*, C. Xia*, J. L. Close, M. Zhang, J. He, Z. Huang, A. R. Halpern, B. Long, J. A. Miller, E. S. Lein, X. Zhuang#. Conservation and divergence of cortical cell organization in human and mouse revealed by MERFISH. Science 377, 56-62 (2022). [Full text]
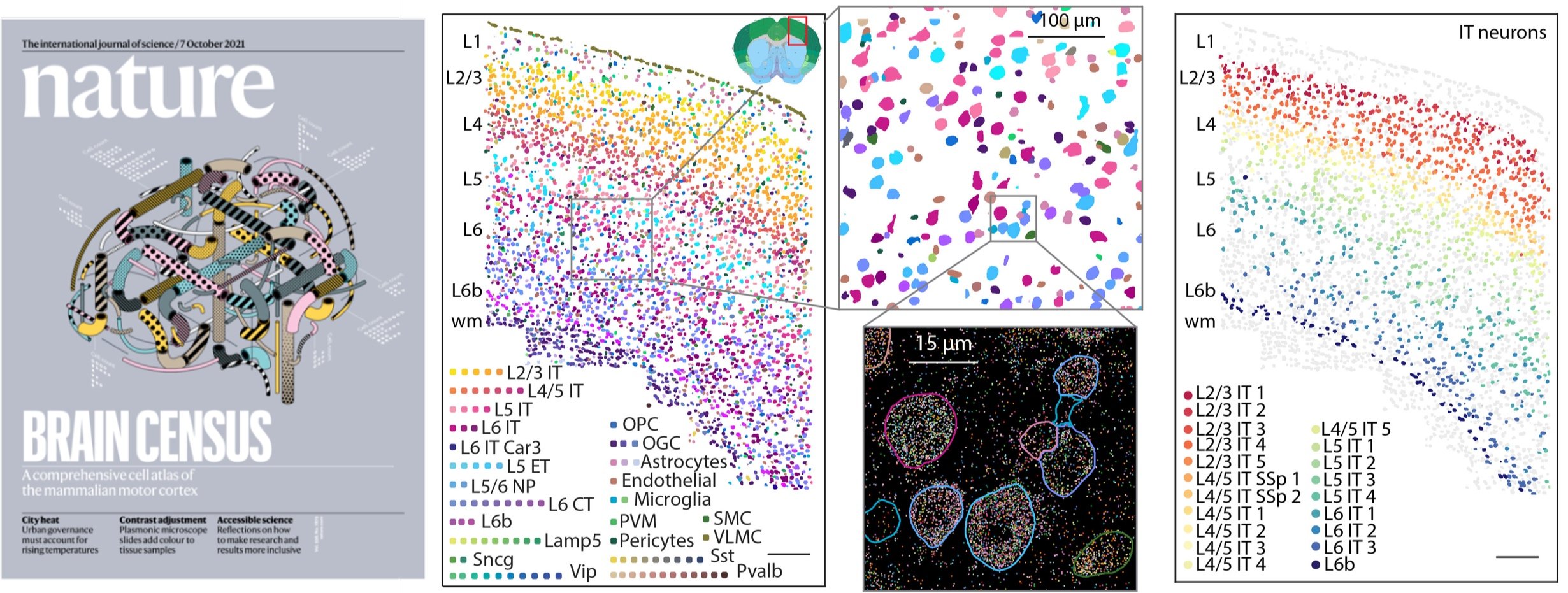
M. Zhang, S. W. Eichhorn, B. Zingg, Z. Yao, K. Cotter, H. Zeng, H. Dong, X. Zhuang#. Spatially resolved cell atlas of the mouse primary motor cortex by MERFISH. Nature 598, 137–143 (2021). [Full text]
T. Biancalani#, G. Scalia, L. Buffoni, R. Avasthi, Z. Lu, A. Sanger, N. Tokcan, C. R. Vanderburg, Å. Segerstolpe, M. Zhang, I. Avraham-Davidi, S. Vickovic, M. Nitzan S. Ma, A. Subramanian, M. Lipinski, J. Buenrostro, N. B. Brown, D. Fanelli, X. Zhuang, E. Z. Macosko, A. Regev#. Deep learning and alignment of spatially resolved single-cell transcriptomes with Tangram. Nat Methods 18, 1352–1362 (2021). [Full text]
BRAIN Initiative Cell Census Network (BICCN). A multimodal cell census and atlas of the mammalian primary motor cortex. Nature 598, 86–102 (2021). [Full text]
D. Wang, Y. Qin, M. Zhang, X. Li, L. Wang, X. Yang, D. Zhong#. The origin of ultrafast multiphasic dynamics in photoisomerization of bacteriophytochrome. J. Phys. Chem 11, 5913-5919 (2020).
M. Zhang, L. Wang, D. Zhong#. (Invited Review) Photolyase: Dynamics and electron-transfer mechanisms of DNA repair. Arch. Biochem. Biophys. 632, 158-174 (2017).
M. Zhang, L. Wang, D. Zhong#. (Invited Review) Photolyase: Dynamics and mechanisms of repair of sun-induced DNA damage. Photochem. Photobiol. 93, 78-92 (2017).
M. Zhang, L. Wang, S. Shu, A. Sancar, D. Zhong#. Bifurcating electron-transfer pathways in DNA photolyases determine the repair quantum yield. Science 354, 209-213 (2016). [Full text]
J. Gao*, X. Wang*, M. Zhang*, M. Bian*, W. Deng, Z. Zuo, Z. Yang#, D. Zhong#, C. Lin#. Trp triad-dependent rapid photoreduction is not required for the function of Arabidopsis CRY1. Proc. Natl. Acad. Sci. U.S.A. 112, 9135-9140 (2015).
Z. Liu, M. Zhang, X. Guo, C. Tan, J. Li, L. Wang, A. Sancar#, D. Zhong#. Dynamic determination of the functional state in photolyase and the implication for cryptochrome. Proc. Natl. Acad. Sci. U.S.A. 110, 12972-12977 (2013).
M. Simon, J. A. North, J. C. Shimko, R. A. Forties, M. B. Ferdinand, M. Manohar, M. Zhang, R. Fishel, J. J. Ottesen#, M. G. Poirier#. Histone fold modifications control nucleosome unwrapping and disassembly. Proc. Natl. Acad. Sci. U.S.A. 108, 12711-12716 (2011).
M. Ma, S. Chatterjee, M. Zhang, D. Bong#. Stabilization of vesicular and supported membranes by glycolipid oxime polymers. Chem Commun (Camb) 47, 2853-2855 (2011).